Listeria monocytogenes is an ubiquitous Gram-positive bacterium responsible for listeriosis. Worldwide, this foodborne disease is relatively rare but serious. About 1,500 cases per year are recorded in Europe and 2,500 per year in US. Because of its high fatality rate (between 20 and 30 %), listeriosis ranks among the most frequent cause of human death due to foodborne illnesses (EFSA-ECDC, 2014).
L. monocytogenes causes severe symptoms such as septicemia and meningitis for the immunocompromised and elderly population as well as for pregnant women giving birth to stillborn infants or infected new born children (Goulet et al. 2012). The clinical feature in healthy people is mainly described as asymptomatic carriage or mild febrile gastroenteritis (Ooi et al. 2005).
Listeriosis mainly affects a specific group of the population with increased susceptibility, the YOPI (young, old, pregnant and immunosuppressed) (Farber and Peterkin, 1991). Since these populations, mainly elderly, are in constant increase in our countries, the control of L. monocytogenes is an important issue. In case of infection, it is difficult to identify the source since L. monocytogenes has a long latency period of 1 to 91 days (Goulet et al., 2013). Listeriosis can take three different forms with regard to the host: i) gastrointestinal listeriosis, ii) systemic listeriosis and iii) abortion and neonatal listeriosis.
The main route of transmission to humans is through consumption of contaminated food. L. monocytogenes can be found in raw foods and in processed foods that are contaminated during manufacturing, post-processing or storage.
Known host species
L. monocytogenes can infect animals and humans.
Samples used for detection of L. monocytogenes
Food and human samples
Nomenclature
The genus Listeria belongs to the Firmicute phylum, the Bacilli class, the Bacilliales, order and the Listeriaceae family. L. monocytogenes contains 13 distinct somatic (O) antigenic types (i.e. 1/2a, 1/2b, 1/2c, 3a, 3b, 3c, 4a, 4b, 4c, 4d, 4e, 4ab and 7) and 5 flagellar (H)-antigenic types (A, B, C, D and E). L. monocytogenes is divided into four lineages: I, II, III and IV (Orsi et al., 2011). The serotypes involved in human listeriosis are mainly 1/2a and 4b, representing ca. 90% followed by 1/2b ca. 8% and, to a lesser extent, 1/2c and 3a ((EFSA-ECDC, 2014)). Serotypes 1/2a, 1/2b and 1/2c are frequently isolated from food processing environments (Orsi et al., 2011).
Recent typing studies indicate that a few important clonal complexes account for a majority of listeriosis outbreaks and sporadic cases in Europe and worldwide (Ragon et al., 2008; Chenal-Francisque et al., 2011) (Cantinelli et al., 2013).
Each clonal complex gathers several sequence types separated from one central sequence type by one allelic mutation, as described by Ragon et al. (2008).
Genome
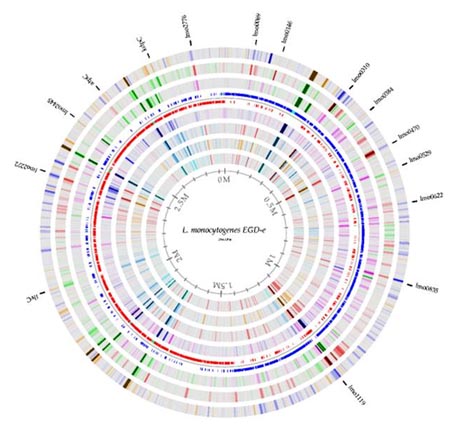
Figure 1. The EGD-e genome created in GenomeAtlas available here: http://www.cbs.dtu.dk/services/GenomeAtlas-2.0/
L. monocytogenes has a chromosome with a size of about 3 Mbp. The EGD-e reference strain genome is of 2 944 528 bp with 39% GC (Bécavin et al., 2014).
To date, L. monocytogenes is divided between 82 clonal complexes and 183 Sequence Types not related to a clonal complex as defined by the Pasteur Institute (http://bigsdb.web.pasteur.fr/perl/bigsdb/bigsdb.pl?db=pubmlst_listeria_seqdef_public&page=browse).
To date, NCBI’s RefSeq database contains 55 circular L. monocytogenes genomes.
Compare reference set
Last update: August 2015
The L. monocytogenes reference set consists of 18 publically available genomes, which represents genomes of the most important Clonal Complexes: CC1, CC2, CC3, CC4, CC5, CC6, CC7, CC8, CC9, CC11, CC14, CC69, CC121, CC131, CC155, CC361 and two genomes not included in any CC representing ST551 and ST617.
References
Becavin, C., Bouchier, C., Lechat, P., Archambaud, C., Creno, S., Gouin, E., Wu, Z., Kuhbacher, A., Brisse, S., Pucciarelli, M.G., Garcia-Del Portillo, F., Hain, T., Portnoy, D.A., Chakraborty, T., Lecuit, M., Pizarro-Cerda, J., Moszer, I., Bierne, H., and Cossart, P. (2014). Comparison of widely used Listeria monocytogenes strains EGD, 10403S, and EGD-e highlights genomic variations underlying differences in pathogenicity. MBio 5, e00969-00914. doi: 10.1128/mBio.00969-14.
Cantinelli, T., Chenal-Francisque, V., Diancourt, L., Frezal, L., Leclercq, A., Wirth, T., Lecuit, M., and Brisse, S. (2013). "Epidemic clones" of Listeria monocytogenes are widespread and ancient clonal groups. J Clin Microbiol 51, 3770-3779. doi: 10.1128/JCM.01874-13.
Chenal-Francisque, V., Lopez, J., Cantinelli, T., Caro, V., Tran, C., Leclercq, A., Lecuit, M., and Brisse, S. (2011). Worldwide distribution of major clones of Listeria monocytogenes. Emerg Infect Dis 17, 1110-1112. doi: 10.3201/eid/1706.101778.
Efsa-Ecdc (2014). The European Union Summary Report on Trends and Sources of Zoonoses, Zoonotic Agents and Food-borne Outbreaks in 2012. EFSA Journal 12, 3547-3312. doi: doi:10.2903/j.efsa.2014.3547
Farber, J.M., and Peterkin, P.I. (1991). Listeria monocytogenes, a food-borne pathogen [published erratum appears in Microbiol Rev 1991 Dec;55(4):752]. Microbiol Rev 55, 476-511.
Goulet, V., King, L.A., Vaillant, V., and De Valk, H. (2013). What is the incubation period for listeriosis? BMC Infect Dis 13, 11. doi: 10.1186/1471-2334-13-11.
Orsi, R.H., Den Bakker, H.C., and Wiedmann, M. (2011). Listeria monocytogenes lineages: Genomics, evolution, ecology, and phenotypic characteristics. Int J Med Microbiol 301, 79-96. doi: 10.1016/j.ijmm.2010.05.002.
Ragon, M., Wirth, T., Hollandt, F., Lavenir, R., Lecuit, M., Le Monnier, A., and Brisse, S. (2008). A new perspective on Listeria monocytogenes evolution. PLoS Pathog 4, e1000146. doi: 10.1371/journal.ppat.1000146.